Large-scale genetic study sheds light on the causes of hemorrhoids
International research team identified pathophysiological mechanisms of hemorrhoidal disease by studying the DNA of nearly one million people.
Found at the very end of the digestive tract, hemorrhoids are blood-filled cushions that help contain stool and control defecation in humans. Often confusing, the word hemorrhoids is also used to refer to hemorrhoidal disease (or piles), in which hemorrhoids enlarge, causing itching, burning, eczema, soiling and bleeding, thus limiting everyday activity. While the symptoms mostly remain mild enough to be resolved with conservative treatment (usually grade I-II hemorrhoids), more severe forms of hemorrhoidal disease require surgical treatment associated with considerable disease burden (usually grade III-IV hemorrhoids). They affect a large part of the population, while the causes are mostly unknown. Many risk factors have been suggested including a sedentary lifestyle, obesity, reduced dietary fiber intake, spending excess time on the toilet or straining during defecation and strenuous lifting, with several being controversially discussed. Possibly due to their intimate nature, hemorrhoidal disease has been generally understudied, and neither the exact molecular mechanisms nor the reasons why only some people develop advanced disease are known, hence undocumented theories, myths and hypotheses have become manifest even in the scientific literature. Now new research published in the journal Gut provides important insights, through world-wide collaboration and the analysis of the genetic make-up of almost one million people.
In the new study, coordinated by Professor Andre Franke of the Institute of Clinical Molecular Biology (IKMB) at Kiel University, Germany, and Professor Mauro D’Amato of the research institute CIC bioGUNE in Bilbao, Spain, researchers looked at millions of DNA changes in the genome of 218,920 patients and 725,213 healthy people from UK Biobank, 23andMe and other population-based cohorts. They identified 102 regions in the human genome that contain genes contributing to increased risk of hemorrhoidal disease.
Further they performed single cell analyses from tissue samples of hemorrhoids. Thus, they could show, that the newly discovered risk genes are primarily expressed in blood vessels and gastrointestinal tissues, and collectively involved in controlling smooth muscle function and the development and integrity of epithelial and endothelial structures in the gut. “Our findings indicate that hemorrhoidal disease results, at least in part, from dysfunction of smooth muscle, epithelial and connective tissue. This points to specific mechanisms in an otherwise poorly characterized common condition, and may lead to the development of additional non-invasive therapeutic options in the long run” commented Andre Franke, who is also a member of the German Cluster of Excellence “Precision Medicine in Chronic Inflammation” (PMI).
The researchers also summarized the new genetic information into polygenic risk scores (PRS) and tested them on three other cohorts, that were not included in the original analysis. Indeed, higher PRS were associated with increased risk of hemorrhoidal disease, younger age at diagnosis, and need for surgery in the analysed 180,435 additional individuals. The team also reported genetic similarities between hemorrhoidal disease and several other gastrointestinal, psychiatric and cardiovascular conditions, which could be verified as most common comorbidities in further analyses of 8 million people from the Danish National Patient Registry.
“Our PRS results require further study in additional cohorts and diverse ethnic groups, but may later help to identify people at increased risk of developing hemorrhoidal disease who could be more closely monitored and to a greater degree benefit from preventative lifestyles”, said Mauro D’Amato.
In addition to the IKMB and CIC bioGUNE, researchers and clinicians from several other institutions participated in the study, including 23andMe Inc, the Mayo Clinic and the University of Michigan from USA, Monash University in Australia, the Karolinska Institute in Sweden, the Institute of Genomics, University of Tartu from Estonia, the University of Copenhagen in Denmark, the University Medical Center Rostock in Germany, Proctological Office Kiel, and others.
Scientific contacts:
Prof. Dr. Andre Franke
Institute of Clinical Molecular Biology (IKMB)
Kiel University (CAU) and University Medical Center Schleswig-Holstein (UKSH)
+49 431 500-15110
a.franke@ikmb.uni-kiel.de
Prof Mauro D’Amato
Gastrointestinal Genetics Lab
CIC bioGUNE – BRTA
mdamato@cicbiogune.es
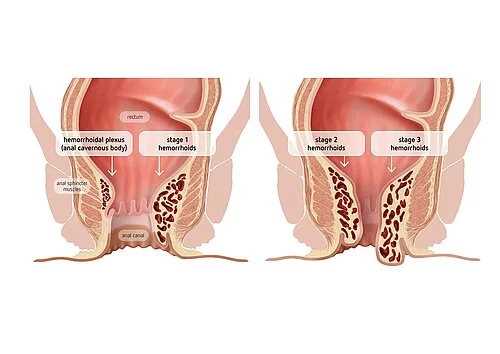
Hemorrhoids are blood-filled arteriovenous “cushions” located in the anal canal and mainly serve as a fine closure thereby contributing to fecal continence. Enlarged hemorrhoids (stage I-IV) can become symptomatic and are then referred to as hemorrhoidal disease that is very common in the population. Advanced symptomatic stages of hemorrhoids often require surgical intervention.
About one million DNA samples were analyzed using modern biochips (so-called SNP arrays).
Original publication:
Zheng T, Ellinghaus D, Juzenas S, et al.: Genome-wide analysis of 944,133 individuals provides insights into the etiology of hemorrhoidal disease. Gut (2021). DOI: 10.1136/gutjnl-2020-323868
About the Cluster of Excellence PMI
The Cluster of Excellence "Precision Medicine in Chronic Inflammation" (PMI) is being funded from 2019 to 2025 through the German Excellence Strategy (ExStra). It succeeds the "Inflammation at Interfaces” Cluster, which was already funded in two periods of the Excellence Initiative (2007-2018). Around 300 members from eight institutions at four locations are involved: Kiel (Kiel University, University Medical Center Schleswig-Holstein (UKSH), Muthesius University of Fine Arts and Design, Kiel Institute for the World Economy (IfW), Leibniz Institute for Science and Mathematics Education (IPN)), Lübeck (University of Lübeck, University Medical Center Schleswig-Holstein (UKSH)), Plön (Max Planck Institute for Evolutionary Biology) and Borstel (Research Center Borstel - Leibniz Lung Center).
The goal is to translate interdisciplinary research findings on chronic inflammatory diseases of barrier organs to healthcare more intensively, as well as to fulfil previously unsatisfied needs of the patients. Three points are important in the context of successful treatment, and are therefore at the heart of PMI research: the early detection of chronic inflammatory diseases, the prediction of disease progression and complications, and the prediction of individual responses to treatment.
Press office
fbuhse@uv.uni-kiel.de+49 (0)431/880 4682 https://precisionmedicine.de
Cluster of Excellence "Precision Medicine in Chronic Inflammation"
Scientific Office
Head: Dr. habil. Susanne Holstein Postal
Christian-Albrechts-Platz 4, 24118 Kiel, Germany
Contact: Sonja Petermann
+49 (0)431 880-4850, fax: +49 (0)431 880-4894
spetermann@uv.uni-kiel.de
Twitter: PMI @medinflame